PROTECT update
Manchester / Rajesh
Department of Data Science Methods, Julius Center, University Medical Center Utrecht
2025-07-13
Updates in PROTECT methods
swap conditional hazard ratio with marginal hazard ratio
calculations for model checks
- correct marginalization incorporating prior likelihood of latent factor
- compare against cross-validated baseline likelihoods (not full-batch)
Conditional vs marginal hazard ratio
- realization: PROTECT models a conditional hazard ratio for treatment (conditional on latent factor, … some other variables)
- trials calculate a marginal hazard ratio
- these are incommensurable
- can calculate a marginal hazard ratio by simulating survival times from hypothetical trials using inferred parameters and patient data by setting treatment to 0 and 1 for every patient
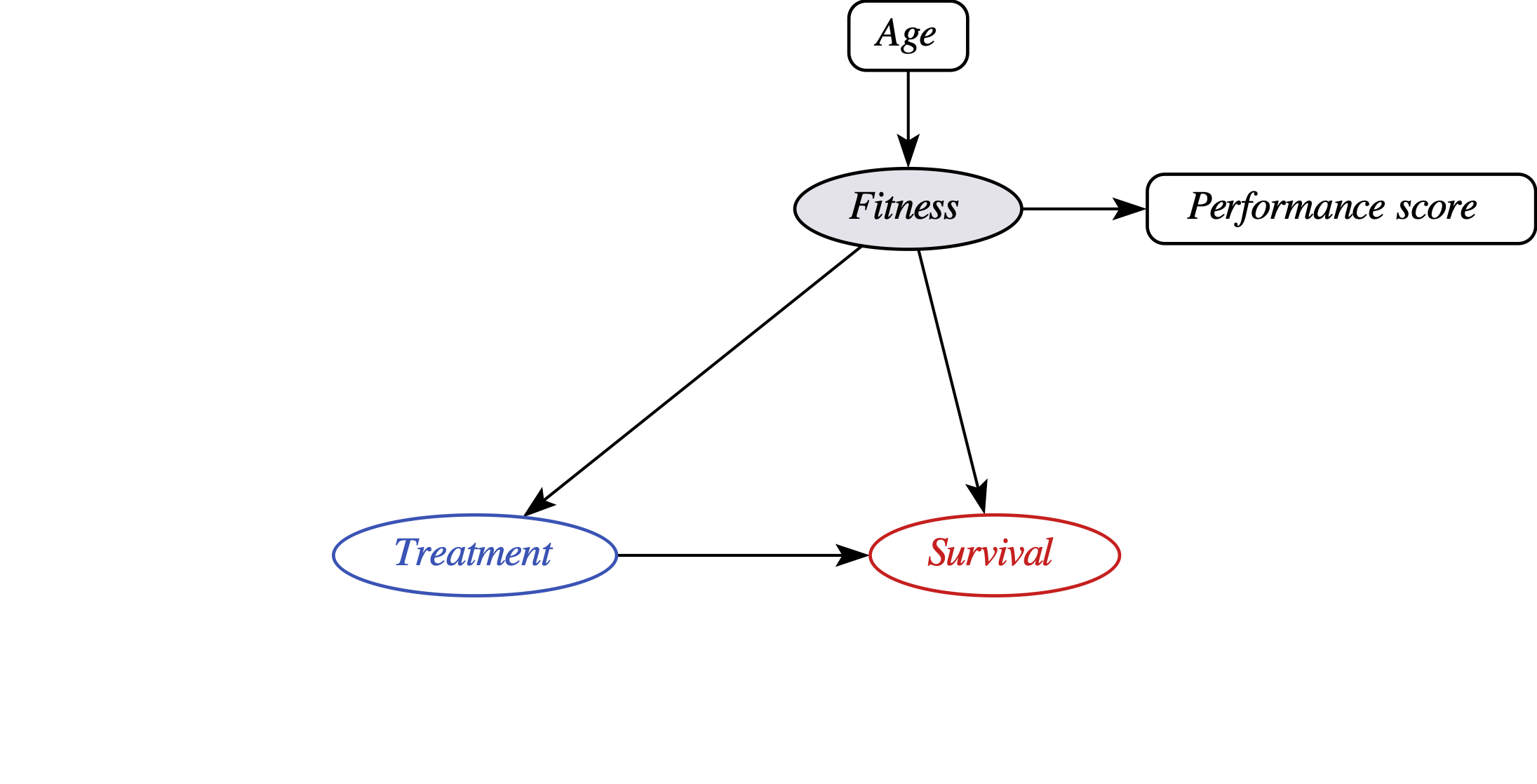
PROTECT uses a latent factor model for the unobserved confounder
- PROTECT models the joint likelihood of ‘observables’:
- treatment \(T\), survival \(Y\) and proxies \(W\) of fitness (performance score, fraily)
- … conditional on ‘controls’ \(X\) (age, stage, histology)
- in addition to ‘global parameters’ \(\theta\) that any regression model has (e.g. treatment to outcome regression coefficient), PROTECT has ‘local parameters’ (local to every patient \(i=1, ... N\)): \(F_i\)
PROTECT inference
- by running MCMC on the ‘training data’, we get samples from the posterior distribution over PROTECT’s parameters:
\[\begin{align*} p(\theta,F|T,Y,W,X) &\propto p(Y,T,W|X,\theta,F) p(F|X, \theta) p(\theta) \end{align*}\]
- in PROTECT, the joint likelihood is formulated as the product of the observation sites following the causal factorization:
\[\begin{align*} p(Y,T,W|X,\theta,F) = p(Y|T,X,\theta,F) p(T|X,\theta,F) p(W|X,\theta,F) \end{align*}\]
- during inference, the posterior for latent factor \(F_i\) of patient \(i\) is informed by their full observed data: \(X_i, W_i, T_i, Y_i\)
PROTECT model checks
- multiple models may be considered compatible with prior knowledge (in practice: different sets of priors)
- in Utrecht, we found for some priors PROTECT copied the treatment into the latent factor, leading to a violation of (a form of) positivity, and unidentifiedness of the treatment effect
- PROTECT has data-driven model checks to assess whether the latent factor ‘effectively communicates’ information between all its dependents (W, T, Y)
- for this, we need distributions over \(F\) using partial information (i.e. different ‘posterior predictive modes’), e.g. ‘no-\(Y\)’:
\[P(F_i|W_i,T_i,X_i,\theta = \theta_s)\]
- note that for every sample \(\theta_s\) of the posterior distribution of global parameters, this distirbution is different
Nested integratoin
\[P(F_i|W_i,T_i,X_i,\theta = \theta_s)\]
- to calculate predictive likelihoods, for example for \(Y\) given \(X,W,T\), we need to do nested integration:
\[\begin{align*} l(Y_i|X_i,W_i,T_i,\{\theta_s\}) &= \sum_s l(Y_i|X_i,W_i,T_i,\theta_s) \\ &= \sum_s \int_{-\infty}^{\infty}l(Y_i|X_i,W_i,T_i,f,\theta_s)p(f|T_i,W_i,X_i,\theta_s)df \end{align*}\]
Nested integration done right
- we need to numerically approximate the integrals
- this approximation should be unbiased and good enough to not be sensitive to e.g. the random seed for MCMC
- we can approximate by running separate MCMC for every sample \(\theta_s\), or a subset
- because this is effectively a large set of 1D integrals, can use faster approximation
approximation in original PROTECT
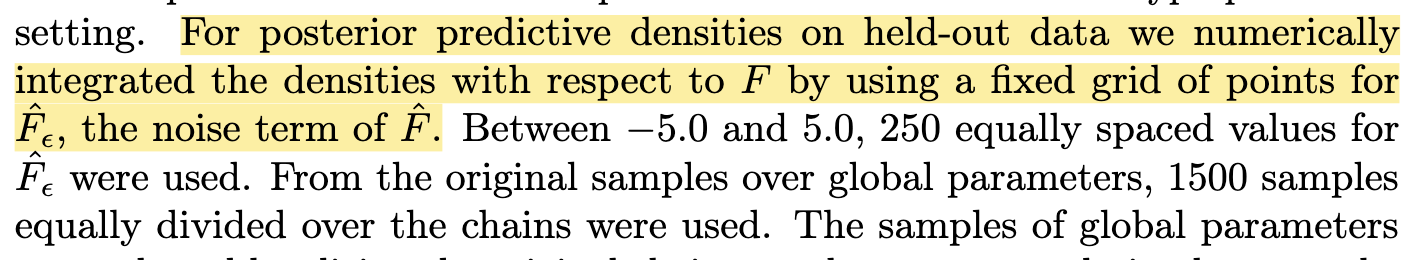
- take a grid of values for \(f\), for each value (and a given \(\theta_s\)), calculate the joint likelihood of the observables for that posterior predictive mode (e.g. \(W_i,T_i\))
- calculate the weighted sum of the log-likelihood of observed \(Y_i\), weighted by the joint likelihood of the \(F-\)conditioning observables
- omission: calculation did not weigh in the prior probability of \(f\), so the predictive likelihoods do not correspond to the actual PROTECT model
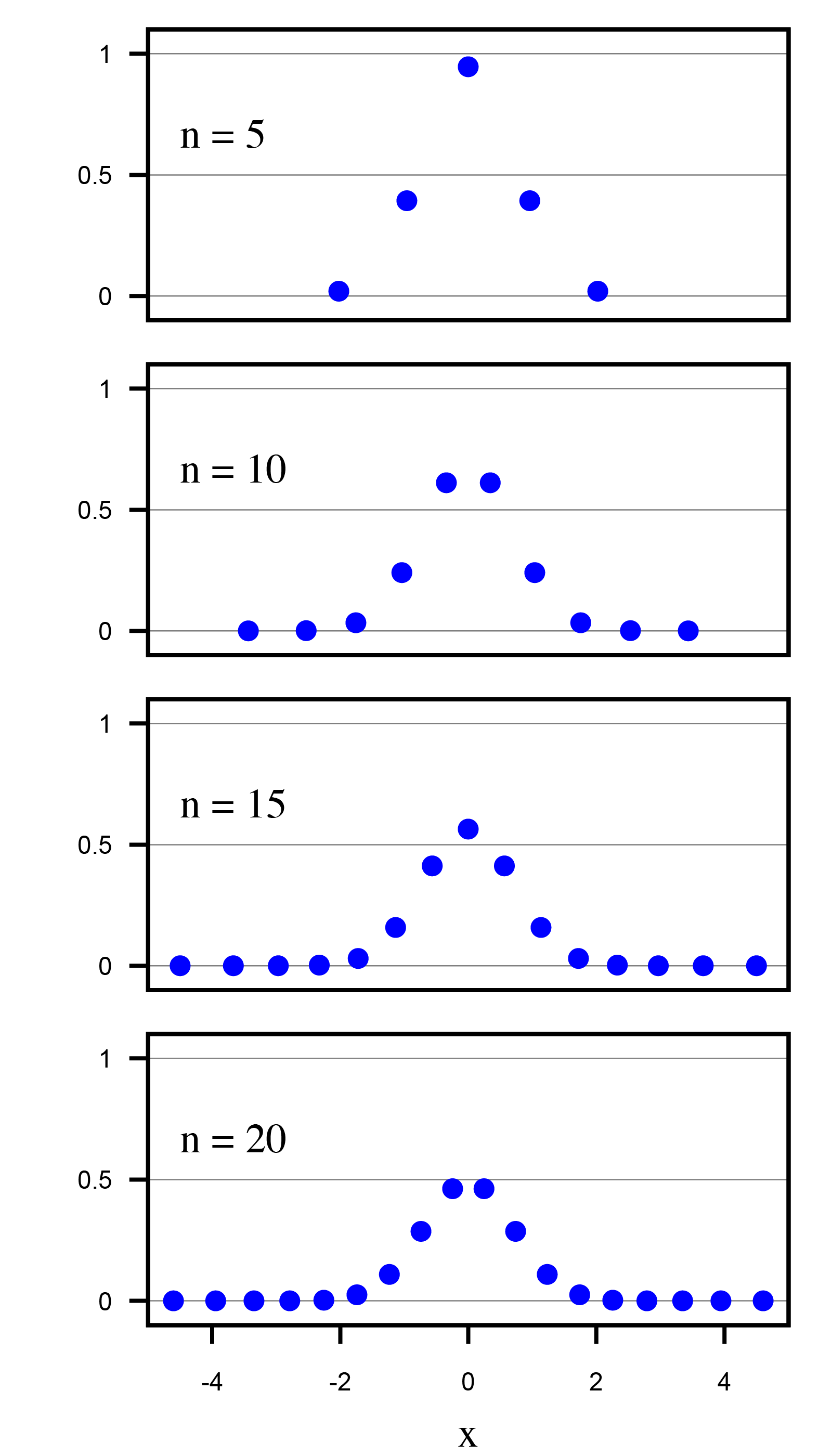
‘new’ solution: use Gauss-Hermite Quadrature
- to integrate over a gaussian distribution, pick \(K\) carefully chosen points \(X_k\) with corresponding weights \(W_k\)
- implemented in PROTECT library, consequences:
- with much fewer points (\(K=32\)),
- much better approximation,
- incorporates prior over \(f\)
- the new approach is tested in cases with analytically knwon likelihoods (gaussian outcome and proxies)
New Utrecht results
- rct: log(HR) -0.17
- before:
- selected hyperparameters: s_bftx: 2.5, s_bfy: {0.1, 1.0, 2.5}
- b_txbinary_y_marginal 0.010 (sd: 0.223, 95 hdi: -0.409 - 0.450)
- now:
- selected hyperparameters:
- sbftxs = np.array([0.1, 2.5, 10.0, 10.0, 100.0, 100.0])
- sbfys = np.array([1.0, 1.0, 1.0, 2.5, 1.0, 2.5])
- b_tx_y_marginal: 0.067 (sd: 0.229, 95 hdi: -0.356 - 0.525)
- marginalized HR: 0.033 (sd: 0.20)
- selected hyperparameters:
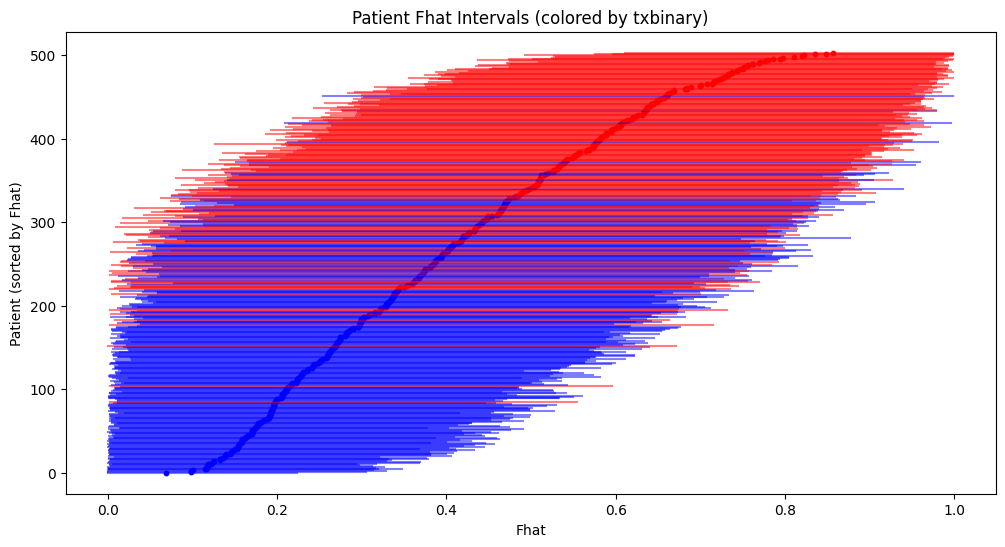
Utrecht overlap?
©Wouter van Amsterdam — WvanAmsterdam — wvanamsterdam.com/talks